1. Why is the circular RNA fire? Where is it sacred? The birth of two articles [Nature[1][2] in 2013 completely overturned our traditional understanding of RNA and quickly detonated the entire biomedical world! After a rigorous statistical summary, the total number of projects related to circular RNA research in the National Natural Science Foundation approved in 2017 was 176, including two outstanding youth funds, one outstanding youth fund, two key projects, 94 items. Projects on the face, 62 youth fund projects, 15 regional science fund projects.


From the perspective of the approved disciplines , it is found that the direction of oncology is the most disciplined subject in the approved projects of circular RNA research, and the digestive system-related tumors are the most approved subjects in the discipline. In addition, the circulation system is ring-shaped. RNA research is also a sudden emergence, becoming the second largest research field of circular RNA. In fact, in addition to oncology and circulatory systems, the medical direction has also received funding in other directions, such as Chinese medicine and pharmacology. I believe that other types of tumors and disciplines in the future will certainly seek more funding for circular RNA projects. At the same time, the research on circular RNA in the direction of life sciences is slightly understated. From the 16 approved projects, basic genetics and bioinformatics (C06) and animal husbandry (C1701) have received more attention. I believe that if we break through the technical bottlenecks in related research fields, there will be more circular RNA research projects in the life sciences.

From the analysis of the approved research content , it was found that compared with 2016, only a few topics for the analysis of cyclic RNA expression were performed in the projects approved in 2017, and most of them were aimed at the functional mechanism of circular RNA, indicating that Circular RNA research has enabled excessive research from basic screening to specific functional mechanisms. It is worth mentioning that the interaction between circular RNA and microRNA is the most common at present, and it is also the research idea with the largest proportion of items. In addition, there are many exosome-derived circular RNA projects, which are also the current circulating RNA. Important direction.
Then, Xiaobian will explain the current research ideas of circular RNA from the following aspects.
What is a circular RNA? Circular RNAs (circRNAs) are a class of non-coding RNA molecules that do not have a 5' end cap and a 3' end poly(A) tail and form a ring structure by covalent bonds. The circRNAs currently found are mainly derived from the exon of the gene, but there are other types, such as intron, intergenic, antisense, and overlapping overlapping.
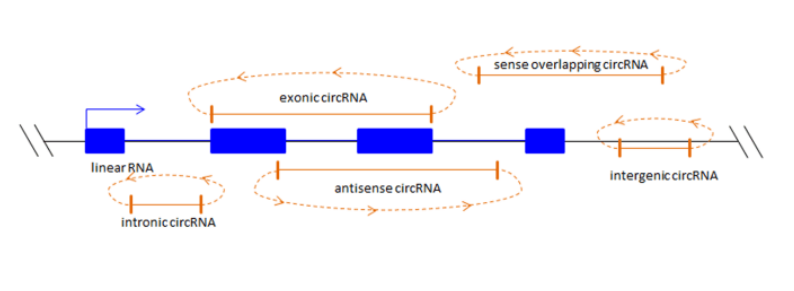
What are the main features of circRNA? (1) Due to trans-shear, circRNA is abundantly present in the
cytoplasm of eukaryotic cells (localization of cytoplasm is a sufficient and necessary condition for the study of miRNA sponge mechanism), and a small number of intron-derived circRNAs exist in nucleic acids and have certain
Tissue specificity, timing and disease specificity (grey is often suitable as a molecular marker). CircRNA is widely present in human cells, sometimes more than 10 times its linear isomer.
(2) Compared with traditional linear RNA, circRNA molecules have no 5' end cap and 3' end poly(A) tail, which is a closed loop structure, which is not easily degraded by exonuclease RNaseR and is more stable than linear RNA. (
RNaseR tolerance test to verify the circular structure of circRNA)
(3) Part of the circRNA contains a miRNA response element, which acts as a competitive endogenous RNA (ceRNA), binds to the miRNA, acts as a
miRNA sponge in the cell, thereby releasing the inhibitory effect of the miRNA on the target gene, and up-regulating the expression level of the target gene. . (Specific research see below)
(4) It can be translated into protein, but most of it is non-coding RNA. (Combined with Appearance: Translating Proteins by M6A Methylation [3])
2. What is the research idea of ​​circular RNA? The so-called soldiers and horses have not moved, the grain and grass first! The study of circular RNA is also a war without smoke, and it is necessary to seize the opportunity to know ourselves and know each other. In order to win in this "war", Yun Xu Bio has prepared a collection of ideas for circular RNA research, as follows.
From the perspective of the workload, difficulty and magazine publishing, Xiaobian divided the research ideas of circular RNA into three categories: expression profiling, biomarkers and functional mechanisms. Then the next series will be detailed from these three perspectives.
Regardless of the angle, it is first necessary to determine the basic information such as the research species, disease model, and sample type. Next, differentially expressed circRNAs were screened by high-throughput means, RNA-seq (the
cloud sequence provides full transcriptome-seq or circRNA-seq ) or reported in the literature. If you want to publish a 3-5-point article in a short time, then
expression profiling seems to be a good fit. If there are a large number of clinical samples, it
is also very easy to make
molecular markers . The ideas and expression profiling are similar to the following figure, but the statistics and verification of clinical samples are more. (
such as KM curve, ROC curve, PAM analysis, etc. )
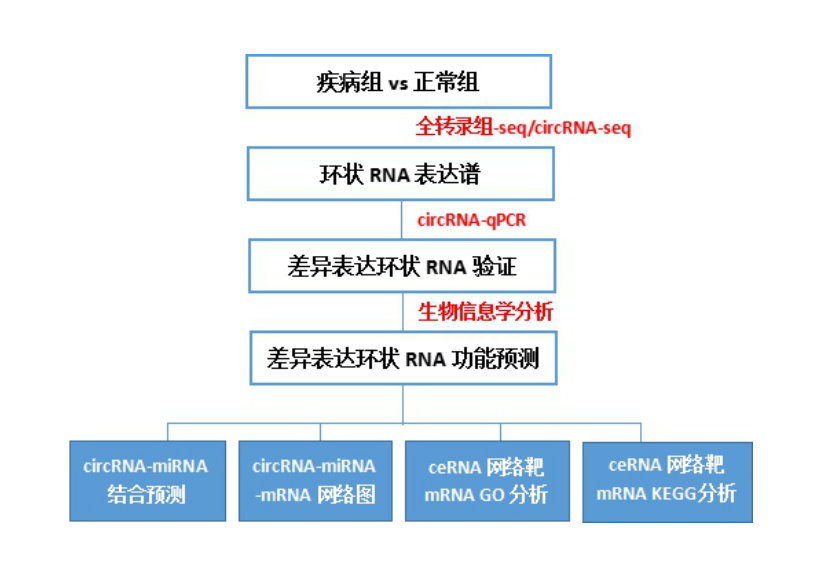
Next, it is the highlight, how to study the function of circRNA function?
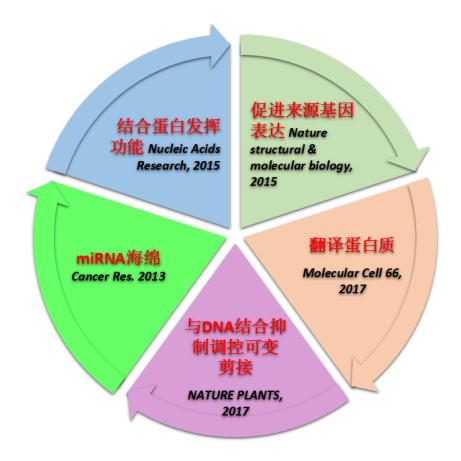
It depends on which angle you want to start with. At present, the functional mechanisms of circRNA research are broadly divided into:
miRNA sponge, binding to functional proteins, homeostatic regulation of source gene expression, and RNA methylation translatable proteins on circRNA . Among them, miRNA sponge is the most common one, and it is also the most natural in the country . The next small series will introduce the research ideas and research methods in this area . miRNA sponge mechanism: After obtaining a batch of data through high-throughput sequencing, how many people will be guilty of how to use the data? Presumably the following article [4] everyone is familiar with it? Then Xiaobian once again took everyone to appreciate the high-profile circular RNA.

The data generated by high-throughput sequencing requires further experimental means for validation. In high-scoring magazines such as Nature communication, verification is all-encompassing. It not only includes expression verification, but also structural verification, cell localization and functional verification. Some even include studies on the mechanism of formation of circular RNA, which are not described here.
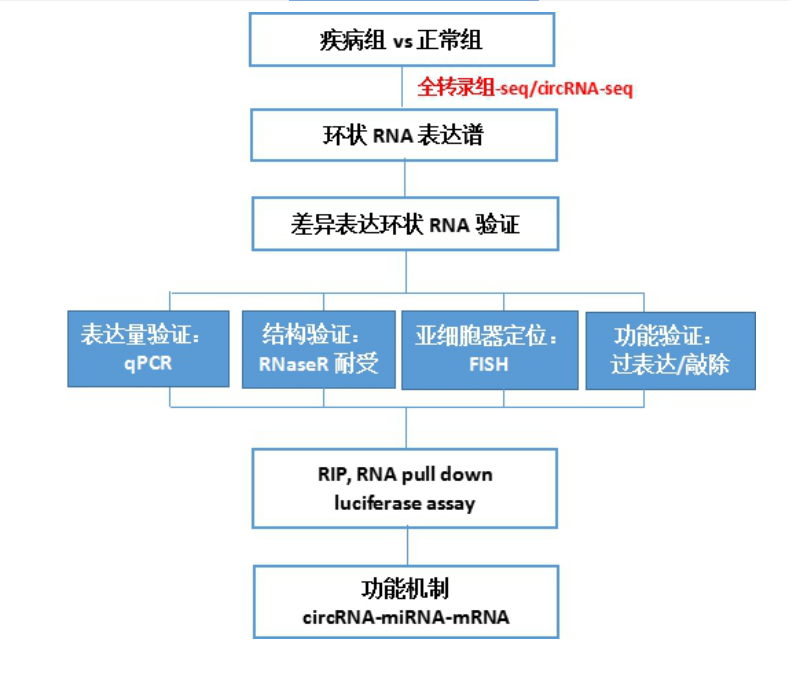
Verification of differential circular RNA in vitro a expression verification: qPCR
b. Structural verification: Since circRNA is not easily degraded by exonuclease RNaseR, it is more stable than linear RNA. The ring structure can be verified by RNase R tolerance experiments.
c cell localization: Since circRNA plays a miRNA sponge mechanism in the cytoplasm, it needs to be verified in the cell by nuclear separation assay or FISH.
d Functional verification: siRNA or overexpression plasmid was designed for 1-2 circRNAs selected from the above steps, and the cell function of circRNA was verified by a series of cell phenotype experiments. Finally, the phenotypes with the most significant phenotypes were selected for follow-up mechanism studies.
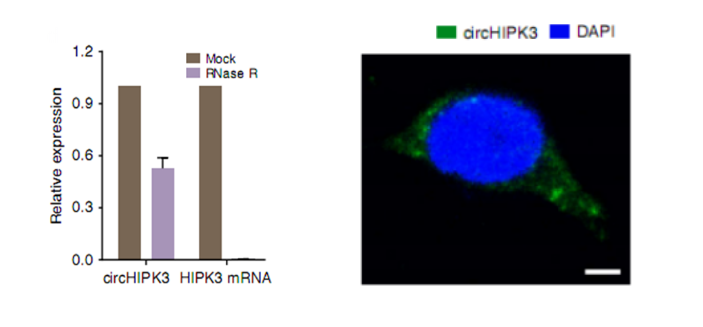
Graphic: Differential circRNA validation left: RNaseR tolerance experiment right: FISH localization
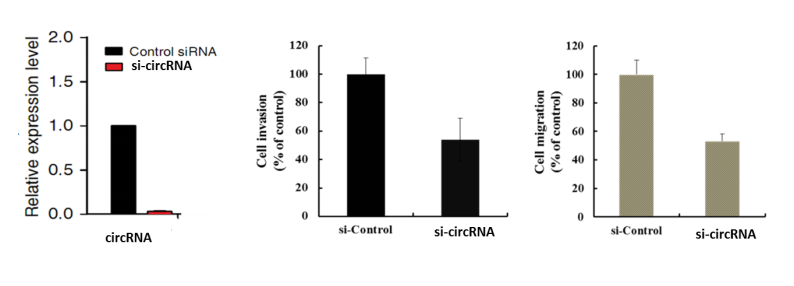
Illustration: Differential loop circRNA left: Knockout experiment: Cell invasion Right: Cell migration assay
Mechanism study of target circRNA a RIP-qPCR: Select one of the most functional circRNAs for RIP-qPCR to detect whether circRNA binds to AGO2 protein. (AGO2 is an indicator protein for circRNA to act as a sponge)
b RNA pull down: The RNA pull down experiment was performed on the above circRNA, and the pulled RNA was subjected to quantitative PCR detection to detect the 10-20 disease miRNA bound to the circRNA in the biosignal prediction.
c luciferase assay: According to the results of the previous step, three or so miRNAs were selected for circRNA for subsequent luciferase assay (miRNA and circRNA expression trends are consistent), and luciferase vector of circRNA and luciferase vector of miRNA binding site mutation were constructed respectively. .
The circRNA luciferase vector was co-transfected into cells with three miRNA mimics to detect fluorescence activity, demonstrating the binding of circular RNA to miRNA mimics.
d miRNA functional assay: The effect of a miRNA on a certain cell phenotype and target gene expression level (eg, migration) was selected;
e Restoration experiments: separate studies: transfection of nc mimics alone or transfection of miRNA mimics alone or transfection of circular RNA overexpression plasmids, and co-transfection of circular RNA + miRNA mimics for a defined cell phenotype and loop The effect of RNA expression levels.
f Replenishment experiment: After knocking out circRNA, the target gene is overexpressed to detect whether the target gene can reverse the phenotypic changes caused by circRNA knockout.
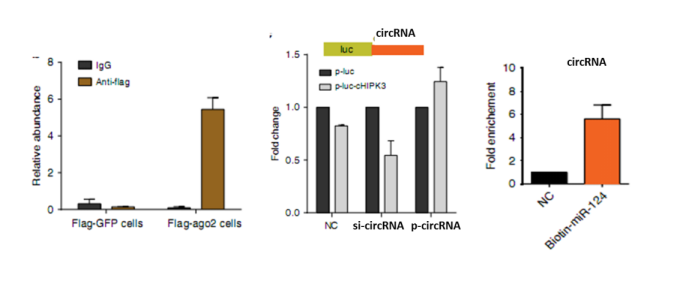
Illustration: Differential circular RNA left: Ago2 RIP: luciferase screening system Right: miRNA pull down
Since it is a true feedback, then Xiaobian here to give you a more easy to get research sponge mechanism ideas as shown below. You can do circRNA-seq and AGO2 RIP-seq at the same time , so that the result of the overlap is the circRNA that may play a sponge role. Furthermore, the interaction between circRNA and AGO2 was reverse verified by RNA pull down-WB, and other functional experiments were consistent with the above.
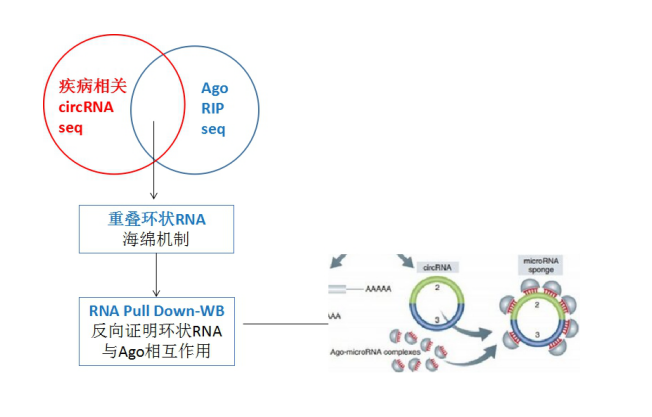
But if my circRNA is not a sponge mechanism, what should I do? Don't be afraid, Xiaobian gives you a big move!
CircRNA binding functional protein: When you find that your circRNA is not playing a sponge, is there a feeling of tears running?
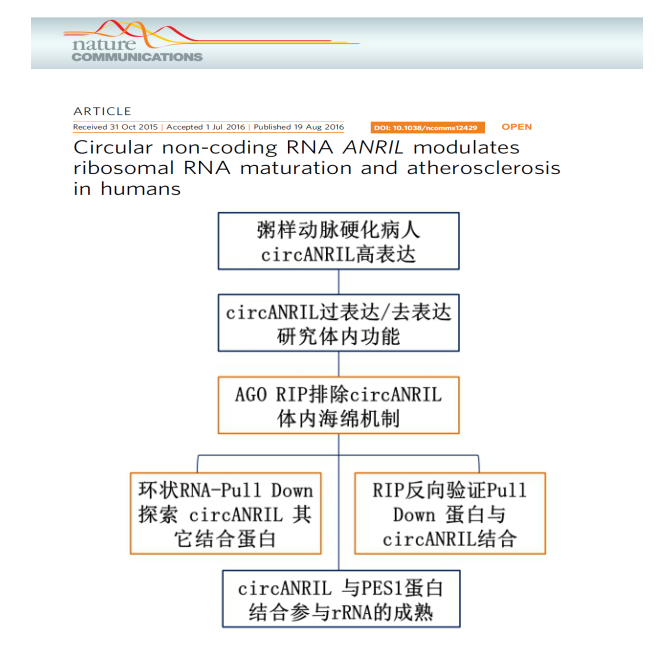
Don't be afraid, Xiaobian will give you a reassurance. It turns out that circRNA can not only play a role in sponge interaction with miRNA, but also function in combination with functional proteins! The following is a good example! Like everyone else, the author of this article first verified whether his circANRIL combined with AGO2 through the AGO2 RIP experiment, but unfortunately, the results confirmed that he thoroughly and miRNA sponge said goodbye, and suddenly Xiaobian also pinched a cold sweat . The author miraculously reversed the situation, explored the binding protein on circANRIL through RNA pull down, and confirmed by RIP that circRNA participates in the maturation process of rRNA by binding to PES1 protein. I want to come to work hard, this article is also successfully published in Nature communication [5].
CircRNA homeostatic regulation of source gene expression: Here, Xiaobian has prepared a circular mechanism that has been hidden for a long time: The research idea of
circRNA homeostatic regulation of source gene expression is to regulate transcription by specific RNA-RNA interaction.

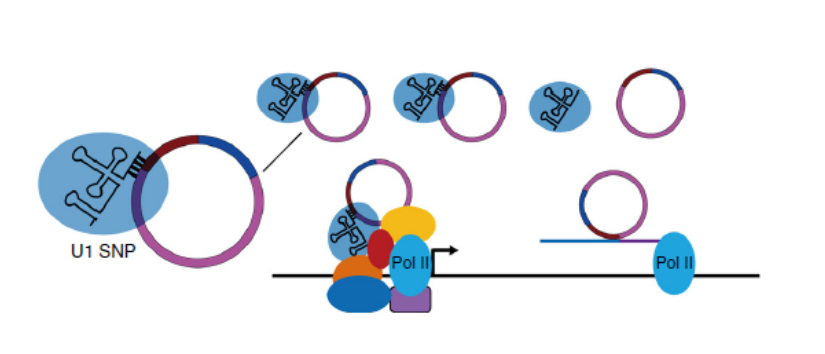
In this paper, the authors first discovered that some circRNAs were bound to RNA polymerase II by CLIP-seq (detecting RNA downstream of RNA-binding proteins). And by identification, it was found that the circular RNA formed by the exon and the intron retained between the exons was named as exon-intron circRNAs (EIciRNAs), in which the significantly enriched circEIF3J and circPAIP were used as follow-up Research object. This was found to be localized to the
nucleus by FISH localization, suggesting that the author may not be able to follow the path of the miRNA sponge.
So what? The authors confirmed the expression of the source gene by silencing circEIF3J and circPAIP by antisense nucleotide technology, and confirmed its regulated gene expression.
RNA-DNA colocalization FISH also confirmed that EIciRNAs co-localize with the source gene in the nucleus. In addition, the interaction between these two EIciRNAs and Pol II also led the author to make a bold conjecture: EIciRNAs may regulate the genes they are derived from.
With this conjecture, the authors followed the
RNA pull down (detection of RNA and RNA/protein action) and CHIRP (a method of detecting DNA-protein interactions with RNA binding). EIciRNAs interact with proteins and RNA. It was found that EIciRNAs not only bind to Pol II but also bind to U1A and U1C snRNP (
which is involved in the processing of RNA in the nucleus of eukaryotic cells. The snRNA and many proteins bind together to become small nuclear ribonucleoproteins, snRNP. Participate in the splicing of messenger RNApre-mRN, making the latter a mature mRNA ( ) and interacting within the nucleus (
FISH ). CHIRP experiments confirmed that EIciRNAs bind to Pol II and U1 snRNP at 300 bp upstream of the transcription start site of the source gene, and binding of U1 snRNP is required [8].
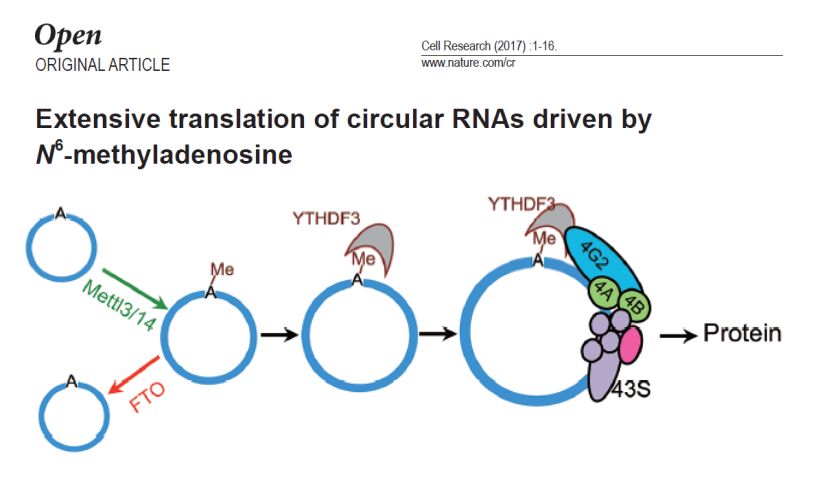
circRNA translation protein: If all this is not enough to attract your attention, then the next case may make you "fall through the glasses." Initially we thought that circRNA is a non-coding RNA and cannot function as a protein. So is this really the case? This article by Cell Research [3] tells you: big mistakes! Cyclic RNA has no free 5' and 3' ends, so if it can be translated it must be through a way that does not depend on the 5' hat structure. Regulatory elements according to the currently reported circular RNA translation proteins include both IRES and m6A modifications.
It is known that some mRNAs can initiate translation from the middle of mRNA by a cis-regulatory element called the internal ribosome entry site (IRSE) independent of the cap structure. Based on this idea, the authors found that they found their own circular RNA. There is no IRES, but the same protein can be translated. However, it has been reported that m6A can regulate translation, but there is no systematic study. The authors boldly suspected that m6A modification occurred on the circular RNA and regulated protein translation.
As expected, the translation of circular RNA can be regulated by RNA methylase. Through a series of experiments, the authors confirmed that YTHDF3 recognizes m6A modifications occurring on circRNA, and recruits translation initiation factors eIF3A and eIF4G2 to initiate translation of proteins.
Not only that, the proteins encoded by circular RNA are still functional, as described in the following two articles [6] [7].

Speaking of the research ideas of translation proteins, it is a little more complicated, so Xiaobian will give you a summary.
1. Translation initiation component predictive analysis:
IRES is a shorthand for the internal ribosome entry site, an RNA regulatory element that has a ribosome recruitment and ribosome assembly and subsequent reading frame translation.
2. Reading frame prediction:
The reading frame is short for Open Reading Frame, which refers to the start of ATG in the nucleic acid sequence (AUG in RNA), three base sets, until TAA, TAG or TGA (A to U in RNA) stop codon sequence.
3. Vector expression verification experiment:
By constructing the normal element and the element mutation vector, the predicted IRES or m6A modification site is mutated to see if the subsequent translation is proceeding normally.
4. Reading frame expression verification experiment:
By adding a tag sequence at the front end of the original reading frame, including a label such as 3×Flag, it is verified whether the reading frame product can be realized under physiological conditions.
5. Identification of endogenous translation products:
Western and other detection experiments were carried out by preparing specific antibodies. This part requires antibody-specific proof experiments, including overexpression and knockout experiments of mock translation products, and specificity can be identified by WB and IP down-spectrum mass spectrometry.
6. Ribosomal binding analysis:
A ribosome separation technique based on sucrose density gradient centrifugation is used to identify whether or not a ribosome binds to a circular RNA, and how much is bound.
7. Translation product overexpression and knockdown (division) experiments:
By constructing an overexpression or knockdown/knockout system, the effects of different levels of translation products on cell physiological indices were observed.
3. Circular RNA project country natural application proposal
(1) Avoid the research content of simple design expression spectrum. Perform expression profiling as early as possible, find the target circular RNA with research value, and carry out some simple verification experiments, which should form the outline of the general functional mechanism research.
(2) Carry out novel research directions of circular RNA. This includes the screening and identification of circular RNAs related to pathological or physiological models that have not been reported or rarely reported, or mechanisms such as miRNA sponges, interactions with functional proteins, binding to apparent levels, and translation of protein orientation.
(3) A variety of research tools for efficiently analyzing the interaction of circular RNA with other molecules or proteins. Including RIP, RNA pull-down, CLIP and other experimental techniques, if you can get valid data, it will add a lot of color in the application of the National Natural Science Foundation of China.
references
1.Memczak S, Jens M, Elefsinioti A, et al. Circular RNAs are a large class of animal RNAs with regulatory potency. [J]. Nature, 2013, 495(7441):333.
2. Hansen TB, Jensen TI, Clausen BH, et al. Natural RNA circles function as efficient microRNA sponges [J]. Nature, 2013, 495(7441):384-388.
3.Yun Y, Fan X, Mao M, et al. Extensive translation of circular RNAs driven by N6-methyladenosine[J]. Cell Research, 2017, 27(5):626.
4. Zheng Q, Bao C, Guo W, et al. Circular RNA profiling reveals an abundant circHIPK3 that regulates cell growth by sponging multiple miRNAs[J]. Nature Communications, 2016, 7(11215):11215.
5.Holdt LM, Anika S, Kristina S, et al. Circular non-coding RNAANRILmodulates ribosomal RNA maturation and atherosclerosis in humans: [J]. Nature Communications, 2016, 7:12429.
6. Legnini I, Timoteo GD, Rossi F, et al. Circ-ZNF609 Is a Circular RNA that Can Be Translated and Functions in Myogenesis[J]. Molecular Cell, 2017, 66(1):22.
7.Yang Y, Gao X, Zhang M, et al. Novel Role of FBXW7 Circular RNA in Repressing Glioma Tumorigenesis. [J]. Journal of the National Cancer Institute, 2018, 110(3).
8.Li Z, Huang C, Bao C, et al. Exon-intron circular RNAs regulate transcription in the nucleus. [J]. Nature Structural & Molecular Biology, 2015, 22(3): 256.
Cloud order biology related recommendation
Whole transcriptome sequencing
Circular RNA sequencing
RIP sequencing
RNA pull down
Circular RNA real-time quantitative PCR

Shanghai Yunxu Biological Technology Co., Ltd.
 Shanghai Cloud-seq Biotech Co.,Ltd
        Address: 6/F, Building 71, Lane 1066, Qinzhou North Road, Caohejing High-tech Development Zone, Shanghai Â
Telephone website:
 mailbox:
Automatic Massage Collapsible Foot Spa Massager
Auto Massage Foldable Foot Spa Massager is a device that provides relaxing and rejuvenating foot massage. It has a collapsible design for easy storage and transport. The massager has multiple massage rollers that target specific pressure points on your feet to deliver a deep tissue massage that helps relieve tension and improve circulation. It also has a heating function that can help soothe sore muscles and promote relaxation. The massager is easy to use and can be controlled with the push of a button. It's a great way to pamper yourself after a long day on your feet or to relieve foot pain and discomfort.
Automatic Massage Collapsible Foot Spa Massager,Folding Foot Bath Machine,Foot Spa,Massager Folding Foot Spa
Huaian Mimir Electric Appliance Co., LTD , https://www.mimirfootbath.com

















